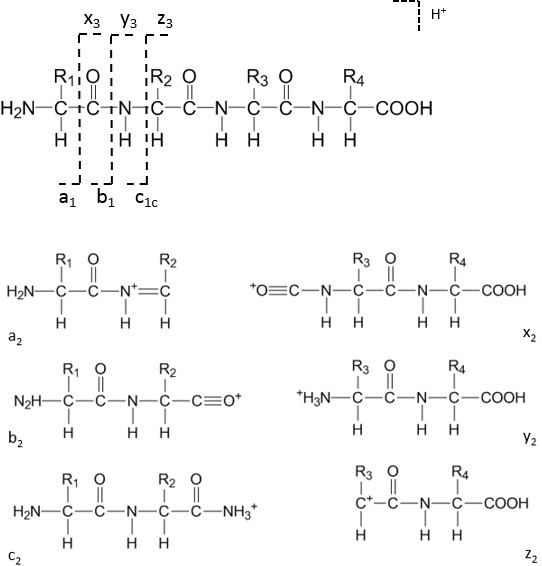
Depending on the fragmentation methods, different types of ions may have quite different intensity values (peak heights), and yet, the ion type information remains unknown from spectrum data. Another challenge is that peptide fragmentation generates multiple types of ions, including a, b, c, x, y, z, internal cleavage, and immonium ions ( 38).

In our de novo sequencing problem, the research is carried to the next extreme, where exactly 1 of 20 L amino acid sequences can be considered as the correct prediction (L is the peptide length, and 20 is the total number of amino acid letters). Furthermore, unlike traditional machine learning methods, those feature layers are not predesigned based on domain-specific knowledge, and hence, they have more flexibility to discover complex structures of the data. The key aspect of deep learning is its ability to learn multiple levels of representation of high-dimensional data through its many layers of neurons. Deep learning has also made its way into biological sciences ( 30). It now forms the core of the artificial intelligence platforms of several technology giants, such as Google, Facebook, and Microsoft, as well as many startups in the industry. Deep learning has recently brought about a revolution in many research fields ( 25), repeatedly breaking state of the art records in image processing ( 26, 27), speech recognition ( 28), and natural language processing ( 29). In this study, we introduce neural networks and deep learning to de novo peptide sequencing and achieve major breakthroughs on this well-studied problem. Not only does our study extend the deep learning revolution to a new field, but it also shows an innovative approach in solving optimization problems by using deep learning and dynamic programming. Moreover, DeepNovo is retrainable to adapt to any sources of data and provides a complete end-to-end training and prediction solution to the de novo sequencing problem. We further used DeepNovo to automatically reconstruct the complete sequences of antibody light and heavy chains of mouse, achieving 97.5–100% coverage and 97.2–99.5% accuracy, without assisting databases. We evaluated the method on a wide variety of species and found that DeepNovo considerably outperformed state of the art methods, achieving 7.7–22.9% higher accuracy at the amino acid level and 38.1–64.0% higher accuracy at the peptide level. The networks are further integrated with local dynamic programming to solve the complex optimization task of de novo sequencing. DeepNovo architecture combines recent advances in convolutional neural networks and recurrent neural networks to learn features of tandem mass spectra, fragment ions, and sequence patterns of peptides. In this study, we propose a deep neural network model, DeepNovo, for de novo peptide sequencing. De novo peptide sequencing from tandem MS data is the key technology in proteomics for the characterization of proteins, especially for new sequences, such as mAbs.
